Our Technology
Transforming Molecular Diagnostics
EasyScreen™ & 3base™ Technology
Our scientists are pioneers of the bisulphite method used for distinguishing methylated cytosine (Methyl C) residues from unmethylated cytosine (C) residues in DNA.
The technology not only serves to identify methylation patterns in the DNA, it can also be used to simplify DNA and RNA sequences more generally, particularly in lower organisms. In unmethylated nucleic acid, the bisulphite method converts all cytosines (C) ultimately into thymine (T), so that Cs disappear from the sequence altogether, resulting in the conversion of a 4 base sequence into a 3 base sequence of only As, Ts and Gs. We refer to this resultant DNA or RNA as ‘3base™’.
The 3base™ process doubles the amount of genetic information we can interrogate for each organism, as the 2 strands of nucleic acid are no longer complementary. We have effectively doubled the genome size and therefore we are able to look for genuinely unique genetic signatures of our target organisms. This extends to viral sequences, even single-stranded viruses, which still produce the complementary strand upon infecting a cell. Again, these 2 strands are no longer complementary post 3base™ conversion, and allows us to identify the different strands at potentially physiologically relevant life-cycle stages.
Our 3base™ technology is more immune than 4 base molecular methods to naturally occurring genetic mutation. This is most beneficial in single-stranded RNA viruses, where a C to T mutation is the most frequent natural change; 3base™ technology obviates this mutation as well as any T to C mutations.
The benefits of 3base™ technology can be summarised as:
Allows for simple multiplexed assays
- Fewer primers, less competition and a more efficient reaction translating to improved sensitivity
- More targets can be detected per specimen, eg 22 gastroenteritis causing enteric pathogens are simultaneously detected from stool
Universally applicable to all specimen types, compatible with both DNA and RNA
- Able to work with multiple specimen types simultaneously
Open platform to suit all testing laboratories
- No need for laboratories to purchase new equipment as 3base™ does not require specialist hardware and is compatible with most automated sample preparation systems and real-time PCR instruments
3 Base Conversion
- Is incorporated into standard sample processing protocols
- By eliminating one-in-four of the naturally occurring bases, 3base™ technology is unique in its ability to simplify the detection of an entire class of microbial species. Only one assay is required for all/any variants within a class, thereby providing a new generation screening methodology (EasyScreen™)
- Sodium bisulphite reacts with cytosine to form uracil
- After amplification all uracil residues are converted to thymine
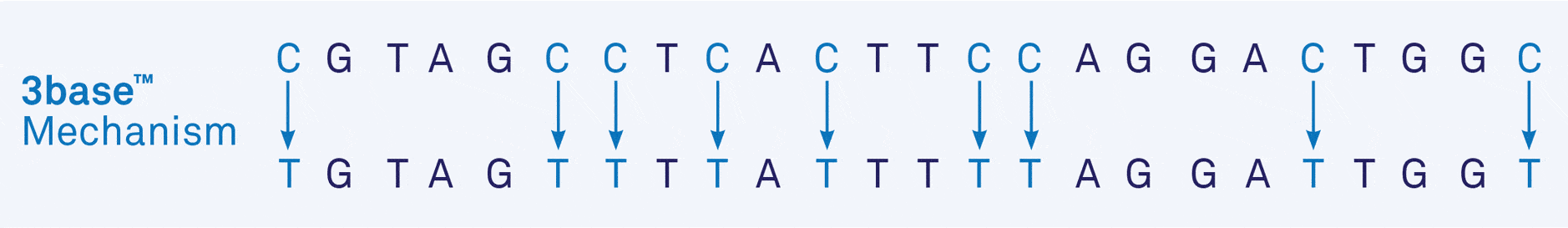
- Figure 1a. Example of the 3base™ mechanism. The example sequences below show the increase in homology from 75% (“Before”) to 95% (“After”) via the 3base™ conversion where all C bases are detected as T bases.
- Significantly improves consensus primer homology across species variants
- Enables the use of single or minimal primers/probes to accurately screen for all/any variants present in a sample
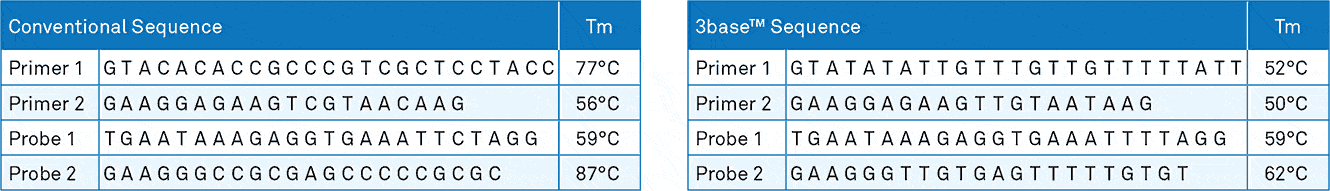
Figure 1b. The regular and 3base™ DNA sequence for two primers and two probes is shown. The primers and probes for the 3base™ have a more similar melting temperature (Tm) improving the efficiency of multiplex real-time PCR.
- The 3base™ converted genome is different from 4-base native genomes; however genomic aspects that identify the presence of disease or microorganisms remain intact
- The converted genome delivers an increase in target homology (similarity) across sub-species
- Genotyping can be effectively carried out using 3base™ genomes
- 3base™ assays will not cross-react with 4-Base native sequences and this reduces the potential of false positive signals due to impurities in commercially available mastermixes or the environment
1 Grunau C, Clark SJ, Rosenthal A (2001). Bisulfite genomic sequencing: systematic investigation of critical experimental parameters. Nucleic Acids Res. Jul 1;29(13): E65-5.
Regulatory:
- EasyScreen™ Pathogen Detection Kits may not be available for in vitro diagnostic use in your jurisdiction. Consult the Pathogen Detection section of this website for more information. MethylEasy Kits are intended for Research Use Only in all jurisdictions.